-Search query
-Search result
Showing 1 - 50 of 136 items for (author: mayer & a)

EMDB-43508: 
Structure of a bacterial gasdermin small oval pore assembly
Method: single particle / : Johnson AG, Mayer ML, Kranzusch PJ

EMDB-43509: 
Structure of a bacterial gasdermin medium oval pore assembly
Method: single particle / : Johnson AG, Mayer ML, Kranzusch PJ

EMDB-43510: 
Structure of a bacterial gasdermin large oval pore assembly
Method: single particle / : Johnson AG, Mayer ML, Kranzusch PJ

EMDB-43511: 
Structure of a bacterial gasdermin double pore assembly
Method: single particle / : Johnson AG, Mayer ML, Kranzusch PJ
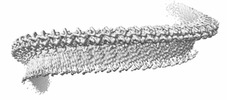

EMDB-43513: 
Structure of a bacterial gasdermin slinky-like oligomer from a heterogeneous sample
Method: single particle / : Johnson AG, Mayer ML, Kranzusch PJ

EMDB-15411: 
Single particle structure of Atg18-WT
Method: single particle / : Mann D, Fromm S, Martinez-Sanchez A, Gopaldass N, Mayer A, Sachse C

PDB-8afx: 
Single particle structure of Atg18-WT
Method: single particle / : Mann D, Fromm S, Martinez-Sanchez A, Gopaldass N, Mayer A, Sachse C

EMDB-18657: 
PROTAC-mediated complex of KRAS with VHL/Elongin-B/Elongin-C/Cullin-2/Rbx1
Method: single particle / : Fischer G, Peter D, Arce-Solano S

EMDB-41983: 
Structure of the phage immune evasion protein Gad1 bound to the Gabija GajAB complex
Method: single particle / : Antine SP, Johnson AG, Mooney SE, Mayer ML, Kranzsuch PJ

PDB-8u7i: 
Structure of the phage immune evasion protein Gad1 bound to the Gabija GajAB complex
Method: single particle / : Antine SP, Johnson AG, Mooney SE, Mayer ML, Kranzsuch PJ

EMDB-15408: 
Tube assembly of Atg18-PR72AA
Method: helical / : Mann D, Fromm S, Martinez-Sanchez A, Gopaldass N, Mayer A, Sachse C

EMDB-15410: 
Tube assembly of Atg18-WT
Method: helical / : Mann D, Fromm S, Martinez-Sanchez A, Gopaldass N, Mayer A, Sachse C

EMDB-15412: 
Subtomogram average of membrane-bound Atg18 oligomers
Method: subtomogram averaging / : Mann D, Fromm S, Martinez-Sanchez A, Gopaldass N, Mayer A, Sachse C

PDB-8afq: 
Tube assembly of Atg18-PR72AA
Method: helical / : Mann D, Fromm S, Martinez-Sanchez A, Gopaldass N, Mayer A, Sachse C

PDB-8afw: 
Tube assembly of Atg18-WT
Method: helical / : Mann D, Fromm S, Martinez-Sanchez A, Gopaldass N, Mayer A, Sachse C

PDB-8afy: 
Subtomogram average of membrane-bound Atg18 oligomers
Method: subtomogram averaging / : Mann D, Fromm S, Martinez-Sanchez A, Gopaldass N, Mayer A, Sachse C

EMDB-17015: 
Cryo-EM map of the focused refinement of the subfamily III haloalkane dehalogenase from Haloferax mediterranei dimer forming hexameric assembly.
Method: single particle / : Polak M, Novacek J, Chmelova K, Marek M

PDB-8ooh: 
Cryo-EM map of the focused refinement of the subfamily III haloalkane dehalogenase from Haloferax mediterranei dimer forming hexameric assembly.
Method: single particle / : Polak M, Novacek J, Chmelova K, Marek M

EMDB-16998: 
Cryo-EM structure of subfamily III haloalkane dehalogenase DhmeA from Haloferax mediterranei
Method: single particle / : Marek M, Novacek J, Polak M, Chmelova K

EMDB-27205: 
Closed state of SARS-CoV-2 BA.2 variant spike protein
Method: single particle / : Zhang J, Tang WC, Gao HL, Shi W, Peng HQ, Volloch SR, Xiao TS, Chen B

EMDB-27206: 
One RBD-up state of SARS-CoV-2 BA.2 variant spike protein
Method: single particle / : Zhang J, Tang WC, Gao HL, Shi W, Peng HQ, Volloch SR, Xiao TS, Chen B

EMDB-27207: 
Middle state of SARS-CoV-2 BA.2 variant spike protein
Method: single particle / : Zhang J, Tang WC, Gao HL, Shi W, Peng HQ, Volloch SR, Xiao TS, Chen B

PDB-8d55: 
Closed state of SARS-CoV-2 BA.2 variant spike protein
Method: single particle / : Zhang J, Tang WC, Gao HL, Shi W, Peng HQ, Volloch SR, Xiao TS, Chen B

PDB-8d56: 
One RBD-up state of SARS-CoV-2 BA.2 variant spike protein
Method: single particle / : Zhang J, Tang WC, Gao HL, Shi W, Peng HQ, Volloch SR, Xiao TS, Chen B

PDB-8d5a: 
Middle state of SARS-CoV-2 BA.2 variant spike protein
Method: single particle / : Zhang J, Tang WC, Gao HL, Shi W, Peng HQ, Volloch SR, Xiao TS, Chen B

EMDB-40570: 
Structure of a bacterial gasdermin slinky-like oligomer
Method: single particle / : Johnson AG, Mayer ML, Kranzusch PJ

PDB-8sl0: 
Structure of a bacterial gasdermin slinky-like oligomer
Method: single particle / : Johnson AG, Mayer ML, Kranzusch PJ

EMDB-29016: 
Cryo-EM structure of SARS-CoV-2 postfusion spike in membrane
Method: single particle / : Zhang J, Shi W, Cai YF, Zhu HS, Peng HQ, Voyer J, Volloch SR, Cao H, Mayer ML, Song KK, Xu C, Lu JM, Chen B
EMDB-29017: 
Cryo-EM structure of SARS-CoV-2 postfusion spike in membrane
Method: single particle / : Zhang J, Shi W, Cai YF, Zhu HS, Peng HQ, Voyer J, Volloch SR, Cao H, Mayer ML, Song KK, Xu C, Lu JM, Chen B

EMDB-29018: 
Cryo-EM structure of SARS-CoV-2 postfusion spike in membrane
Method: single particle / : Zhang J, Shi W, Cai YF, Zhu HS, Peng HQ, Voyer J, Volloch SR, Cao H, Mayer ML, Song KK, Xu C, Lu JM, Chen B

PDB-8fdw: 
Cryo-EM structure of SARS-CoV-2 postfusion spike in membrane
Method: single particle / : Zhang J, Shi W, Cai YF, Zhu HS, Peng HQ, Voyer J, Volloch SR, Cao H, Mayer ML, Song KK, Xu C, Lu JM, Chen B

EMDB-25419: 
Previously uncharacterized rectangular bacteria in the dolphin mouth
Method: electron tomography / : Dudek NK, Galaz-Montoya JG, Shi H, Mayer M, Danita C, Celis AI, Wu GH, Behr B, Huang KC, Chiu W, Relman DA

EMDB-35208: 
Cryo-EM structure of the polyphosphate polymerase VTC complex(Vtc4/Vtc3/Vtc1)
Method: single particle / : Mayer A, Wu S, Ye S

PDB-8i6v: 
Cryo-EM structure of the polyphosphate polymerase VTC complex(Vtc4/Vtc3/Vtc1)
Method: single particle / : Mayer A, Wu S, Ye S

EMDB-29323: 
Structure of RdrA from Escherichia coli RADAR defense system
Method: single particle / : Duncan-Lowey B, Johnson AG, Rawson S, Mayer ML, Kranzusch PJ

EMDB-29324: 
Map of RdrA from Escherichia coli RADAR defense system in single-split conformation
Method: single particle / : Duncan-Lowey B, Johnson AG, Rawson S, Mayer ML, Kranzusch PJ

EMDB-29325: 
Map of RdrA from Escherichia coli RADAR defense system in double-split conformation
Method: single particle / : Duncan-Lowey B, Johnson AG, Rawson S, Mayer ML, Kranzusch PJ

EMDB-29326: 
Structure of RdrA from Streptococcus suis RADAR defense system
Method: single particle / : Duncan-Lowey B, Johnson AG, Rawson S, Mayer ML, Kranzusch PJ

EMDB-29327: 
Structure of RdrB from Escherichia coli RADAR defense system
Method: single particle / : Duncan-Lowey B, Johnson AG, Rawson S, Mayer ML, Kranzusch PJ

EMDB-29328: 
Structure of RdrA-RdrB complex from Escherichia coli RADAR defense system
Method: single particle / : Duncan-Lowey B, Johnson AG, Rawson S, Mayer ML, Kranzusch PJ

PDB-8fnt: 
Structure of RdrA from Escherichia coli RADAR defense system
Method: single particle / : Duncan-Lowey B, Johnson AG, Rawson S, Mayer ML, Kranzusch PJ

PDB-8fnu: 
Structure of RdrA from Streptococcus suis RADAR defense system
Method: single particle / : Duncan-Lowey B, Johnson AG, Rawson S, Mayer ML, Kranzusch PJ

PDB-8fnv: 
Structure of RdrB from Escherichia coli RADAR defense system
Method: single particle / : Duncan-Lowey B, Johnson AG, Rawson S, Mayer ML, Kranzusch PJ

PDB-8fnw: 
Structure of RdrA-RdrB complex from Escherichia coli RADAR defense system
Method: single particle / : Duncan-Lowey B, Johnson AG, Rawson S, Mayer ML, Kranzusch PJ

EMDB-27704: 
Cryo-EM structure of insulin receptor (IR) bound with S597 peptide
Method: single particle / : Park J, Li J, Mayer JP, Ball KA, Wu JY, Hall C, Accili D, Stowell MHB, Bai XC, Choi E

EMDB-27705: 
Cryo-EM structure of insulin receptor (IR) bound with S597 component 2
Method: single particle / : Park J, Li J, Mayer JP, Ball KA, Wu JY, Hall C, Accili D, Stowell MHB, Bai XC, Choi E

PDB-8dtl: 
Cryo-EM structure of insulin receptor (IR) bound with S597 peptide
Method: single particle / : Park J, Li J, Mayer JP, Ball KA, Wu JY, Hall C, Accili D, Stowell MHB, Bai XC, Choi E

PDB-8dtm: 
Cryo-EM structure of insulin receptor (IR) bound with S597 component 2
Method: single particle / : Park J, Li J, Mayer JP, Ball KA, Wu JY, Hall C, Accili D, Stowell MHB, Bai XC, Choi E

EMDB-31642: 
Local construction of SARS-CoV-2 S protein RBD in complex with XG014 Fab
Method: single particle / : Wang K, Wang XX

EMDB-25188: 
Full-length insulin receptor bound with site 1 binding deficient mutant insulin (A-V3E)
Method: single particle / : Bai XC, Choi E
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
